Automated design of genomic Southern blot probes
Mike DR Croning, David G Fricker, Noboru H Komiyama and Seth GN Grant
Introduction
A novel software pipeline for designing and optimizing Southern blot probes in silico for use against genomic DNA targets is described.
The software was written and validated for two reasons:
- To address our own needs to regularly design Southern blot probes, automating this process, reducing the time needed to do this manually.
- To optimize the resultant probes by employing a brute-force search approach to significantly improve the chances of finding the (near) best probe for the loci of interest, the aim being to reduce both the time and expense in the laboratory that results from failed Southern blot assays, and subsequent rounds of probe redesign.
The in silico scoring measures that we developed for evaluating the automated probe designs suggest they should perform as well, or better, than previous manual designs, while reducing the time taken by the molecular biologist to yield a successful probe as we had planned.
We went on to experimentally test around 15 probes in the study and we report this experimental validation in the manuscript. The majority of the probes we tested in Southern blotting performed well confirming our in silico prediction methodology and the general usefulness of the software for automated genomic Southern blot probe design.
The software is freely-available under the terms of the Artistic License 2.0, and we hope that it finds extensive reuse by investigators in the genomics, genetics and molecular biology communities.
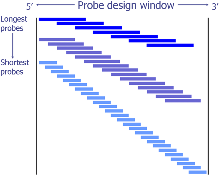
Tiling algorithm for Southern blot probe design
Given user-supplied chromosomal coordinates, and a desirable size range for the southern blot probe, we used a tiling approach to generate many possible probes in the specified design window. The program starts from the maximum allowable probe length, tiling the window by moving by a small percentage of the probe length each time (default 5%). Once this is completed the probe length is reduced by 50 bases (configurable) and the window re-tiled generating more candidate probes. The process is repeated until the minimum probe length is reached.
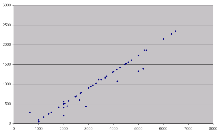
This approach produces a linear relationship between the numbers of candidate probes to search against the target genome with respect to the length of the input design window. With a desirable probe length range of 500-1300bp this produces approximately 900 probes to search for a 3kb input window.
Calibration with 8 experimentally-validated probes
We calibrated the method using a set of 8 manually-designed mouse genomic probes (download here) that we have previously successfully employed for Southern blotting. We searched these against the NCBI m33 genome assembly (see below).
| Probe Name | Gene Target | Length bases | Self / second hit score ratio | Second hit identity (%) | Second hit query coverage (%) | Min repetitive & low-complexity DNA (%) |
|---|---|---|---|---|---|---|
| Dusp6_5prime_probe | Dusp6 (5') | 946 | 30.1 | 71 | 8 | 3.2 |
| SAP102_5prime_PDZ3_probe | Dlg3 (5' PDZ3) | 969 | 27.2 | 72 | 8 | 2.7 |
| Dusp6_3prime_probe | Dusp6 (3') | 1004 | 29.4 | 61 | 13 | 4.5 |
| actb_probe | Actb | 881 | 22.8 | 91 | 6 | 6.7 |
| SAP102_3prime_probe | Dlg3 (3') | 886 | 22.2 | 77 | 8 | 19.4 |
| NR2B_probe | Grin2b | 567 | 11.1 | 81 | 14 | 9.5 |
| SAP102_5prime_probe | Dlg3 | 784 | 9.89 | 68 | 29 | 81.7 |
| PSD-95_exon_9_probe | Dlg4 (exon 9) | 296 | 3.3 | 76 | 54 | nd |
| Average ± standard error | 791.6 ± 85.9 | 19.5 ± 3.6 | 74.6 ± 3.2 | 17.5 ± 5.8 | 18.2 ± 10.8 | |
As can be seen above these had an average length of approximately 800 bp. When searched with Exonerate (with parameters --model affine:local --score 150) all of these produced a perfect match to their genomic locus (as would be expected) and a number of additional lower-scoring alignments to other loci. These second best matches spanned 17.5 ± 5.8% (mean ± standard error) of the probe length, with 74.6 ± 3.2% DNA sequence identity (n=8). From the scores of the 'self' and the highest scoring off-target locus alignments we calculated a score ratio as measure of uniqueness of the candidate probe. Our calibration probes averaged 19.5 ± 3.6. This score ratio is proportional to both the length and sequence identity of the matches. nd=not determined.
Comparing the probe sequences to a version of the genomic assembly that has been screened for repeats and low-complexity regions by RepeatMasker and DUST allows us to estimate the repetitive DNA content of individual probes. Our calibration probes contained 18.2 ± 10.8% such DNA (see Supplementary x).
We chose a minimum score ratio of 10 and a maximum combined repetitive and low-complexity base content of 5% as the minimum requirements for probe acceptance. candidate probes reaching these criteria that were completely overlapped by a longer and better scoring probe are considered redundant and removed from the passing set.
124 automated Southern blot designs
| Probe Design | Mouse Chr | Passed | Genomic Design Window (bases) | Length Best Probe (bases) | Best Score Ratio | Unique Probes | Non-Unique Probes Passed | Total Probes Passed | Total Candidate Probs | Total Probes Passed (%) | Candidate Probes / Kilobase |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | Pass | 4587 | 800 | na | 48 | 8 | 56 | 1559 | 3.6 | 339.9 |
| 2 | 2 | Pass | 6327 | 600 | na | 15 | 461 | 476 | 2277 | 20.9 | 359.9 |
| 3 | 11 | Pass | 2001 | 650 | na | 10 | 127 | 137 | 205 | 66.8 | 102.4 |
| 4 | 11 | Fail | 2001 | 500 | 1.1 | 0 | 0 | 0 | 205 | 0 | 102.4 |
| 5 | 8 | Fail | 1001 | 550 | na | 5 | 0 | 0 | 51 | 0 | 50.9 |
| 6 | 8 | Pass | 1001 | 550 | na | 3 | 48 | 51 | 51 | 100 | 50.9 |
| 7 | 15 | Fail | 1001 | 500 | 10.7 | 0 | 0 | 0 | 51 | 0 | 50.9 |
| 8 | 15 | Pass | 1001 | 500 | na | 22 | 0 | 22 | 51 | 43.1 | 50.9 |
| 9 | 2 | Pass | 2893 | 700 | na | 46 | 6 | 52 | 434 | 12 | 150 |
| 10 | 2 | Fail | 1069 | 550 | 6.2 | 0 | 0 | 0 | 60 | 0 | 56.1 |
| 11 | X | Pass | 16430 | 1300 | na | 214 | 288 | 502 | 3207 | 15.7 | 195.1 |
| 12 | 16 | Pass | 647 | 350 | na | 13 | 23 | 36 | 287 | 12.5 | 443.6 |
| 13 | 16 | Pass | 3964 | 900 | na | 109 | 711 | 820 | 1305 | 62.8 | 329.2 |
| 14 | 16 | Pass | 3660 | 1300 | na | 251 | 535 | 786 | 1179 | 66.7 | 322.1 |
| 15 | 16 | Pass | 2460 | 550 | na | 7 | 292 | 299 | 683 | 43.8 | 277.6 |
| 16 | 16 | Pass | 4260 | 900 | na | 156 | 564 | 720 | 1423 | 50.6 | 334 |
| 17 | 16 | Pass | 3661 | 1300 | na | 253 | 533 | 786 | 1159 | 67.8 | 316.6 |
| 18 | 16 | Fail | 1051 | 500 | 14.5 | 0 | 0 | 0 | 112 | 0 | 106.5 |
| 19 | 16 | Pass | 2461 | 550 | na | 8 | 293 | 301 | 684 | 44 | 277.9 |
| 20 | 16 | Pass | 3171 | 700 | na | 49 | 532 | 581 | 974 | 59.7 | 307.2 |
| 21 | 16 | Pass | 3717 | 600 | na | 22 | 174 | 196 | 1200 | 16.3 | 322.8 |
| 22 | 2 | Pass | 2001 | 550 | 17.5 | 0 | 12 | 12 | 494 | 2.4 | 246.9 |
| 23 | 2 | Fail | 2501 | 500 | na | 2 | 0 | 0 | 700 | 0 | 279.9 |
| 24 | 12 | Pass | 2001 | 1000 | 32.9 | nd | nd | 252 | 405 | 62.2 | 202.4 |
| 25 | 12 | Pass | 2001 | 1000 | 32.9 | nd | nd | 217 | 405 | 53.6 | 202.4 |
| 26 | 3 | Pass | 2001 | 1000 | 31.8 | nd | nd | 249 | 405 | 61.5 | 202.4 |
| 27 | 3 | Pass | 2001 | 1000 | 31.4 | nd | nd | 174 | 405 | 43 | 202.4 |
| 28 | 17 | Pass | 2138 | 1000 | 30.7 | nd | nd | 200 | 446 | 44.8 | 208.6 |
| 29 | 17 | Pass | 5190 | 700 | 21.5 | nd | nd | 21 | 1391 | 1.5 | 268 |
| 30 | 5 | Pass | 4157 | 600 | 17.9 | nd | nd | 19 | 1071 | 1.8 | 257.6 |
| 31 | 5 | Pass | 4998 | 1000 | 30.1 | nd | nd | 261 | 1329 | 19.7 | 265.9 |
| 32 | X | Fail | 2432 | 500 | 10.9 | nd | nd | 0 | 537 | 0 | 220.8 |
| 33 | X | Pass | 2633 | 550 | 19.5 | nd | nd | 6 | 598 | 1 | 227.1 |
| 34 | X | Pass | 3399 | 800 | 23.2 | nd | nd | 155 | 1120 | 13.8 | 329.5 |
| 35 | X | Pass | 5232 | 1000 | 29.8 | nd | nd | 332 | 1865 | 17.8 | 356.5 |
| 36 | X | Fail | 1204 | 400 | 4.5 | nd | nd | 0 | 235 | 0 | 195.2 |
| 37 | X | Pass | 18537 | 1000 | 33.3 | nd | nd | 806 | 7251 | 11.1 | 391.2 |
| 38 | X | Pass | 2001 | 600 | 18 | nd | nd | 30 | 507 | 5.9 | 253.4 |
| 39 | X | Pass | 2001 | 950 | 30.7 | nd | nd | 148 | 557 | 26.6 | 278.4 |
| 40 | X | Pass | 2001 | 600 | 19.5 | nd | nd | 30 | 557 | 5.4 | 278.4 |
| 41 | X | Pass | 2001 | 950 | 30.7 | nd | nd | 148 | 557 | 26.6 | 278.4 |
| 42 | X | Pass | 3501 | 1150 | 35.7 | nd | nd | 303 | 1114 | 27.2 | 318.2 |
| 43 | X | Pass | 3001 | 1150 | 34.8 | nd | nd | 214 | 907 | 23.6 | 302.2 |
| 44 | 17 | Pass | 3001 | 1300 | 39.9 | nd | nd | 11 | 907 | 1.2 | 302.2 |
| 45 | 17 | Pass | 3001 | 1050 | 29 | nd | nd | 355 | 907 | 39.1 | 302.2 |
| 46 | 8 | Pass | 2473 | 1300 | 38.7 | nd | nd | 294 | 686 | 42.9 | 277.4 |
| 47 | 8 | Pass | 4712 | 1150 | 33.2 | nd | nd | 210 | 1611 | 13 | 341.9 |
| 48 | 19 | Pass | 4001 | 1300 | 40.9 | nd | nd | 822 | 1318 | 62.4 | 329.4 |
| 49 | 19 | Pass | 2001 | 1250 | 40.9 | nd | nd | 286 | 494 | 57.9 | 246.9 |
| 50 | 6 | Pass | 2501 | 1300 | 42.2 | nd | nd | 384 | 699 | 54.9 | 279.5 |
| 51 | 6 | Pass | 4501 | 850 | 25.9 | nd | nd | 341 | 1526 | 22.3 | 339 |
| 52 | X | Pass | 1801 | 900 | 29 | nd | nd | 86 | 412 | 20.9 | 228.8 |
| 53 | X | Pass | 6001 | 700 | 19.9 | nd | nd | 55 | 2146 | 2.6 | 357.6 |
| 54 | 6 | Pass | 3001 | 1300 | 41.7 | nd | nd | 193 | 907 | 21.3 | 302.3 |
| 55 | 6 | Pass | 1501 | 500 | 11.7 | nd | nd | 1 | 289 | 0.3 | 192.5 |
| 56 | 11 | Pass | 3501 | 1300 | 40.9 | nd | nd | 270 | 1114 | 24.2 | 318.2 |
| 57 | 11 | Pass | 1401 | 1300 | 28 | nd | nd | 196 | 247 | 79.3 | 176 |
| 58 | 10 | Pass | 3001 | 600 | 19.2 | nd | nd | 15 | 907 | 1.7 | 302.2 |
| 59 | 10 | Pass | 2501 | 900 | 27.8 | nd | nd | 290 | 700 | 41.4 | 279.9 |
| 60 | 11 | Pass | 3001 | 750 | na | 303 | 342 | 645 | 907 | 71.1 | 302.2 |
| 61 | 11 | Pass | 3001 | 1300 | na | 629 | 231 | 860 | 907 | 94.8 | 302.9 |
| 62 | 1 | Pass | 2001 | 600 | na | 13 | 209 | 222 | 494 | 44.9 | 246.9 |
| 63 | 1 | Pass | 2201 | 800 | na | 98 | 269 | 367 | 577 | 63.6 | 262.1 |
| 64 | 7 | Pass | 2701 | 1050 | na | 141 | 107 | 248 | 785 | 31.6 | 290.6 |
| 65 | 7 | Pass | 3001 | 900 | na | 108 | 180 | 288 | 907 | 31.8 | 302.2 |
| 66 | 1 | Pass | 1501 | 600 | na | 17 | 135 | 152 | 289 | 52.6 | 192.5 |
| 67 | 1 | Pass | 6001 | 1300 | na | 330 | 241 | 571 | 2146 | 26.6 | 357.6 |
| 68 | 6 | Pass | 2001 | 500 | na | 4 | 7 | 11 | 494 | 2.2 | 246.9 |
| 69 | 6 | Pass | 4001 | 1050 | na | 1 | 127 | 128 | 1318 | 9.7 | 329.4 |
| 70 | X | Fail | 1701 | 600 | 19.1 | nd | nd | 0 | 368 | 0 | 216.3 |
| 71 | X | Pass | 3501 | 1100 | 34 | 1 | 313 | 314 | 1114 | 28.2 | 318.2 |
| 72 | 8 | Pass | 2001 | 1150 | 38.7 | 0 | 209 | 209 | 494 | 42.3 | 246.9 |
| 73 | 8 | Pass | 5001 | 550 | na | 20 | 320 | 340 | 1729 | 19.7 | 345.7 |
| 74 | 4 | Fail | 1501 | 500 | 13.7 | nd | nd | 0 | 289 | 0 | 192.5 |
| 75 | 4 | Fail | 3001 | 500 | na | 2 | 0 | 0 | 907 | 0.2 | 302.2 |
| 76 | X | Pass | 2001 | 800 | na | 47 | 2 | 49 | 494 | 9.9 | 246.9 |
| 77 | X | Pass | 5001 | 800 | na | 117 | 463 | 580 | 1729 | 33.5 | 345.7 |
| 78 | 2 | Fail | 1501 | 550 | na | 4 | 0 | 0 | 289 | 0 | 192.5 |
| 79 | 2 | Pass | 3001 | 750 | na | 69 | 239 | 295 | 907 | 32.5 | 302.2 |
| 80 | 9 | Fail | 1501 | 900 | 23.3 | 0 | 0 | 0 | 289 | 0 | 192.5 |
| 81 | 9 | Pass | 2501 | 500 | 15 | 0 | 140 | 140 | 700 | 20 | 279.9 |
| 82 | 1 | Pass | 2501 | 850 | na | 273 | 219 | 492 | 699 | 70.4 | 279.5 |
| 83 | 1 | Fail | 3501 | 500 | 15.2 | 0 | 0 | 0 | 1114 | 0 | 318.2 |
| 84 | 17 | Pass | 4457 | 700 | na | 198 | 490 | 688 | 1507 | 45.7 | 338.1 |
| 85 | 17 | Pass | 4001 | 1300 | na | 299 | 73 | 372 | 1318 | 28.2 | 329.4 |
| 86 | 3 | Pass | 3501 | 600 | 22.2 | 0 | 29 | 29 | 1114 | 2.6 | 318.2 |
| 87 | 3 | Pass | 2001 | 650 | 22.2 | 0 | 23 | 23 | 494 | 4.7 | 246.9 |
| 88 | 11 | Fail | 3501 | 500 | 17.9 | 0 | 0 | 0 | 1114 | 0 | 318.2 |
| 89 | 11 | Pass | 1201 | 550 | 16.9 | 0 | 4 | 4 | 166 | 2.4 | 138.2 |
| 90 | 5 | Pass | 4131 | 650 | na | 24 | 222 | 246 | 1372 | 17.9 | 332.1 |
| 91 | 5 | Pass | 2501 | 1300 | na | 408 | 292 | 700 | 700 | 100 | 279.9 |
| 92 | 18 | Pass | 2668 | 850 | 25.9 | 0 | 69 | 69 | 768 | 9 | 287.9 |
| 93 | 18 | Pass | 2001 | 550 | 17.3 | 1 | 10 | 11 | 494 | 2.2 | 246.9 |
| 94 | 5 | Fail | 1811 | 600 | na | 16 | 0 | 0 | 414 | 0 | 228.6 |
| 95 | 5 | Pass | 3296 | 650 | na | 45 | 189 | 234 | 1025 | 22.8 | 311 |
| 96 | 14 | Fail | 1001 | 650 | 5.4 | 0 | 0 | 0 | 95 | 0 | 94.9 |
| 97 | 14 | Pass | 4001 | 700 | na | 52 | 103 | 155 | 1318 | 11.8 | 329.4 |
| 98 | 9 | Pass | 1501 | 1300 | na | 289 | 0 | 289 | 289 | 100 | 192.5 |
| 99 | 9 | Pass | 1501 | 850 | na | 59 | 230 | 289 | 289 | 100 | 192.5 |
| 100 | 4 | Pass | 3001 | 550 | na | 1 | 35 | 36 | 907 | 4 | 302.2 |
| 101 | 4 | Pass | 5001 | 500 | na | 48 | 22 | 70 | 1729 | 4 | 345.7 |
| 102 | 11 | Pass | 2046 | 1300 | na | 300 | 212 | 512 | 512 | 100 | 250.2 |
| 103 | 11 | Pass | 5319 | 850 | 25.9 | 0 | 151 | 151 | 1860 | 8.1 | 349.7 |
| 104 | 13 | Pass | 3001 | 1250 | na | 244 | 11 | 255 | 907 | 28.1 | 302.2 |
| 105 | 13 | Fail | 3501 | 850 | 13.9 | 0 | 0 | 0 | 1114 | 0 | 318.2 |
| 106 | 4 | Pass | 3087 | 750 | na | 65 | 876 | 941 | 941 | 100 | 304.8 |
| 107 | 4 | Pass | 6492 | 1150 | na | 503 | 1004 | 1507 | 2346 | 64.2 | 361.4 |
| 108 | 5 | Pass | 3001 | 850 | 25.8 | 0 | 130 | 130 | 907 | 14.3 | 302.2 |
| 109 | 5 | Pass | 2501 | 950 | na | 79 | 424 | 503 | 700 | 71.9 | 279.9 |
| 110 | 14 | Fail | 1201 | 600 | 15.6 | 0 | 0 | 0 | 166 | 0 | 138.2 |
| 111 | 14 | Pass | 3001 | 700 | na | 234 | 342 | 576 | 907 | 63.5 | 302.2 |
| 112 | 6 | Fail | 2201 | 500 | 9.3 | 0 | 0 | 0 | 577 | 0 | 262.2 |
| 113 | 6 | Pass | 2501 | 600 | 17.6 | 0 | 52 | 52 | 700 | 7.4 | 279.9 |
| 114 | 15 | Fail | 1000 | 549 | 9.4 | 0 | 0 | 0 | 74 | 0 | 74 |
| 115 | 15 | Pass | 4501 | 800 | na | 108 | 22 | 130 | 1526 | 8.5 | 339 |
| 116 | 7 | Pass | 3501 | 1300 | na | 228 | 475 | 703 | 1114 | 63.1 | 318.2 |
| 117 | 7 | Pass | 2501 | 500 | na | 1 | 2 | 3 | 700 | 0.4 | 279.9 |
| 118 | 7 | Pass | 5001 | 950 | na | 420 | 121 | 541 | 1729 | 31.3 | 345.7 |
| 119 | 7 | Pass | 2501 | 700 | na | 29 | 25 | 54 | 700 | 7.7 | 279.9 |
| 120 | 15 | Pass | 1001 | 1000 | 19.5 | 0 | 84 | 84 | 95 | 88.4 | 94.9 |
| 121 | 15 | Pass | 4001 | 1050 | na | 135 | 499 | 634 | 1318 | 48.1 | 329.4 |
| 122 | 11 | Pass | 2501 | 650 | na | 24 | 463 | 487 | 700 | 65.3 | 279.9 |
| 123 | 11 | Pass | 1501 | 500 | 14.6 | 0 | 0 | 0 | 289 | 0 | 192.5 |
| 124 | 11 | Pass | 5001 | 900 | na | 111 | 153 | 264 | 1729 | 15.3 | 345.7 |
| SUMMARY | Total passed | Average length | Average length | Average best score ratio | Average unique probes | Average non-unique probes passed | Average total probes passed | Average per design | Average total probes passed (%) | Average probes / kilobase | |
| 103/124 | 3094.8 ± 202.9 | 818.1 ± 25.0 | 23.7 1.3 | 85.2 ± 14.1 | 176.7 ± 23.1 | 240.8 ± 24.2 | 899.6 ± 72.6 | 28.6 ± 2.6 | 263.4 ± 7.2 | ||
Southern Blot package documentation
Index
Downloads & class documentation
- Perl module/class browser
- Download package: southern_blot_design_25_08_09.tar.gz MD5: e8a1191dc99dca75cfee52ff64212f5f, 206582 bytes
A. Copyright & licensing conditions
Scripts, software and documentation copyright 2005-2009 Genes to Cognition Programme (G2C) and Genome Research Limited (GRL).
You may distribute this file/module under the terms of the artistic licence: http://www.perlfoundation.org/artistic_license_2_0
B. Introduction
Southern blotting is an experimental procedure where DNA, from a genomic or other source, is digested with a restriction enzyme and then separated by size using gel electrophoresis. The fragments are transferred from the gel onto a membrane ('blotted') which is then incubated with a labelled single-stranded DNA probe. Such a procedure allows one to locate a particular sequence of DNA within a complex mixture of DNA. From a gene targeting perspective, Southern blotting can be used to detect whether a targeting event has successfully taken place.
Designing a 'good' Southern blot probe for a particular gene or locus involves finding a stretch of DNA sequence at that locus, generally 500-1000bp long, that has the desirable qualities of being unique to that locus, with little or no repetitive DNA content. Molecular biologists tend to design their probes manually, by excising portions of genome sequence from online genome browsers (such as such as Ensembl) and then pasting them into a genome-search site enabling them to check the genome for sequence hits. Ideally a probe sequence should return a single hit to the region it was designed against, with little or no cross-reactivity to other parts of the genome.
If this is not the case, the investigator will likely shift the piece of DNA chosen a short distance away from what they might consider as the optimal site, and search again. Another option might be to shorten the candidate probe sequence, and repeat the search, particularly if was obvious one end of the initial sequence appeared to be lacking the desired specificity, thus giving rise to the extraneous hits.
With it taking quite a few minutes to perform each round of cutting, pasting and genome searching that proves necessary to find an acceptable sequence for a Southern blot probe, one can appreciate that this does not make effective use of a molecular biologists time, and is very unlikely to find an optimal probe.
The design strategy outlined is clearly very amenable to automation using bioinformatics with the added benefit that the number of candidate probes that can be examined during the design process need not be limited to a few (as when carried out manually) but can be increased to hundreds, or thousands, allowing a very fine-grained analysis to be performed, significantly increasing the chances of finding the best, or at least a near-optimal probe for the chosen locus.
C. Implementation
With the number of genome searches to be carried out potentially taking hours for each probe to be designed, the writing of a single programme which would complete the whole task outlined above is not likely to yield a satisfactory solution.
Instead a more elaborate system is required utilising a database to store and retrieve the design information for each probe, and subsequently the results of the many genome searches carried out for candidate probes. These results can then be analysed to find the best probes.
Such a system would also allow more than one computer to be used to carry out the searches, speeding design, and would also permit the user to modify the selection parameters for the probe, without requiring one to re-run the genome searches for a particular probe, should the initial constraints be found to be too stringent at a particular locus.
A MySQL database (12 tables) was designed for this purpose together with a set of Perl data objects and adaptors to allow programmes to write and retrieve from the database. These follow the design paradigm set by the Ensembl genome analysis system, where one creates a set of (DBEntry) classes for the 'business' objects used by the system, partnered by a set of complementary DBSQL classes that hold the cognate SQL necessary for storing and retrieving from the database. Changes to the database schema can the then be made without impact on the DBEntry classes. The naming conventions used by the classes (and data types returned) generally follow those used in the Ensembl core API, see http://www.ensembl.org/info/docs/api/core
Southern blot design package classes
- GeneTargeting
- ::DBEntry
- ::ComponentHit
- ::Conf
- ::Exonerate
- ::DNAProbe
- ::ExternalDB
- ::Hit
- ::Job
- ::Sequence
- ::Xref
- ::DBAdaptor
- ::BaseAdaptor
- ::ConfAdaptor
- ::DBAdaptor
- ::DNAProbeAdaptor
- ::ExternalDBAdaptor
- ::HitAdaptor
- ::JobAdaptor
- ::SequenceAdaptor
- ::SequenceHitAdaptor
- ::XrefAdaptor
- ::Utils - Grab-bag of general utility methods
- ::Config - Configuration file parsing and setup
- ::Counts
- ::GD - Utilities setting colours etc for GD
- ::Exonerate - Custom exonerate parser
- ::HTMLReport - Simple module for html output
- ::Primer3 - Primer-picking wrapping code
- ::DBEntry
Five programmes were written to perform the Southern blot probe design task in its entirety, along with three accessory scripts:
- create_probe_search_db_tables
- Creates the tables constituting the GeneTargeting MySQL database, allowing one to specify the server parameters on the command line
- create_probe_search
- Given the user-specified chromosomal coordinates for the acceptable design window for the Southern blot probe, enumerates all the possible probes in the window, at the chosen granularity, within the size-range chosen for the probe Creates a number of jobs of class GeneTargeting::DBEntry::Job that are stored in the database each one of which is executed by an instance of run_probe_search
- submit_probe_search
- User-executed script to submit a number of jobs to the compute farm to be utilised, wrapping the underying LSF system
- run_probe_search
- The 'runnable' script used by the nodes of the compute farm to fetch candidate probes from the database, search them against the genome (specified in the config file for create_probe_search) and store the results back for later analysis
- analyse_probe_search
- Used to analyse the results from the all the genome searches, determining which of the candidate probes exceed the minimum acceptable criteria for a Southern blot probe. Results are outputting as static html, including a graphic representation of 'unique', 'good' and 'bad' regions in the previously specified probe design window.
- Also picks primers with Primer3 for recovery of the candidate probes.
Accessory scripts
- get_probe_search_cpu_time
- Calculates the total time taken for the execution of all the jobs making up the Southern blot probe search. Does this by parsing the output files written by LSF
- delete_job_results
- Deletes the results from the database of a job - presumably which are erroneous due to a system failure.
- delete_probe_search
- Deletes the whole probe design from the database, when it is no longer needed or possibly the coorinates were specified incorrectly.
D. Performance
Results seem favourable when compared to a number of manually-designed probes (see the paper) that have been used successfully by Genes to Cognition research programme at the Wellcome Trust Sanger Institute.
Experimentally validation has been performed on a set of the probes automatically designed by the software (see the paper).
E. Available documentation
All the scripts making up the Southern blot probe design system contain POD documentation detailing their use and command line parameters. Should the scripts be run with an invalid parameter combination then the documentation is automatically displayed in order to guide the user.
The configuration files for each of the scripts (present in the conf directory of the package) are commented as to the function and meaning of their various sections and options.
The GeneTargeting API modules utilised for Southern blot design are documented with POD which can be displayed with 'perldoc Module_name.pm' or perldoc classname, such as perldoc GeneTargeting::Utils::Exonerate
Other documentation files include
- southern_blot_design/docs/southern_blot_probe_design.txt - this file
- southern_blot_design/docs/example_run.txt
- southern_blot_design/docs/example_run_output/
F. Package directory structure
- GeneTargeting/
- conf/ - example programme config files
- docs/ - documentation
- modules/
- GeneTargeting/
- DBEntry/
- DBSQL/
- Utils/
- GeneTargeting/
- scripts/ - deployed scripts
- run/ - runner scripts started by pipeline
Environment variables
- GeneTargetingConfDir
- full path to the conf directory
- GeneTargetingBaseDir
- full path to the southern_blot_design directory
H. Program configuration files
Each of the five programmes requires a windows-style .ini configuration file. These should be stored in the directory pointed to by the environment variable GeneTargetingConfDir.
Sections of the supplied .ini configuration files group related configuration options, and are commented.
I. Perl modules required
- Name
- Version tested
- Bio
- 1.5.0
- Bio::Ensembl
- branch 32
- Bio::Tools::Run
- 1.4
- Config::IniFiles
- 2.38
- DBI
- 1.32
- GD
- 2.17
Note other modules may be required by the above modules, please see their documentation should "Can't locate Module.pm" errors arise.
J. Other applications required
- Name
- Version tested
- exonerate
- exonerate-1.0.0
- primer3
- primer3_1.0.0
- LSF
- 5.1 from Platform Computing Corp.