Automated design of genomic Southern blot probes
Mike DR Croning, David G Fricker, Noboru H Komiyama and Seth GN Grant
Design Case - ProbeDesign33 - back to list
Summary
./analyse_probe_search_results : Mon Sep 5 11:27:48 2005
Connected to host: host, as user: user Database : southern_blot_designs Fetched probe design: ProbeDesign33 assembly : NCBIM34 bias : 3prime chromosome : X id : 8 strand : 1 ------------- Design window length: 2633bp All jobs for probe design are successful - total: 30 Fetched conf : Exonerate mouse_NCBIM34 (id: 1) Genome file(s) at: /blastdb/Mouse/NCBIM34/softmasked_dusted Fetched 598 putative probes to analyse Selection criteria to rank probes: Minimum score ratio : 10 (i.e. self-hit must score at least 10 times higher than the next best hit) Maximum percent repetitive bases: 5 Favour probes to the 3prime end of design window ------------------------ Putative probes without hits : 0 Putative probes with single_self_hit : 598/598 Putative probes exceeding selection criteria: 6/598 PASSED (GREEN) AND FAILED PUTATIVE PROBES IN THE DESIGN WINDOW --------------------------------------------------------------
Image
Unique (yellow), passed (green) and failed putative probes in the design window
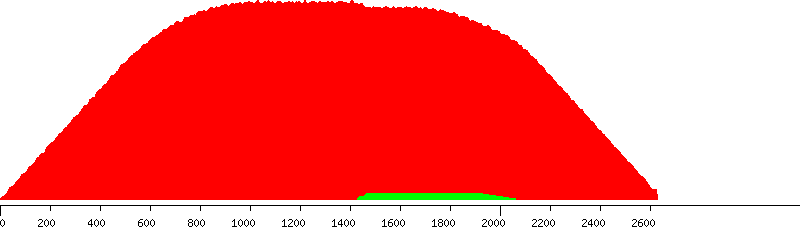
Unique putative probes
No probes
Putative probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 7069 | 550 | 17.9 | 2750 | 1.5 | 621 | 14.9 |
| 6998 | 600 | 19.5 | 3000 | 2.8 | 600 | 13.8 |
| 7151 | 500 | 16.2 | 2500 | 1.6 | 675 | 13.0 |
| 7070 | 550 | 17.9 | 2750 | 2.7 | 648 | 12.4 |
| 6997 | 600 | 15.9 | 3000 | 5.0 | 570 | 5.9 |
Overall score = score_ratio - 2 * repetitive DNA content
Redundant probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 7152 | 500 | 16.2 | 2500 | 3.4 | 700 | 9.4 |
Overall score = score_ratio - 2 * repetitive DNA content
